03:00
3 - What makes a model?
Introduction to tidymodels
Your turn

How do you fit a linear model in R?
How many different ways can you think of?
lmfor linear modelglmnetfor regularized regressionkerasfor regression using TensorFlowstanfor Bayesian regressionsparkfor large data setsbruleefor regression using torch
To specify a model ![]()
- Choose a model
- Specify an engine
- Set the mode
To specify a model ![]()
To specify a model ![]()
- Choose a model
- Specify an engine
- Set the mode
To specify a model ![]()
To specify a model ![]()
To specify a model ![]()
- Choose a model
- Specify an engine
- Set the mode
To specify a model ![]()
To specify a model ![]()
All available models are listed at https://www.tidymodels.org/find/parsnip/
To specify a model ![]()
- Choose a model
- Specify an engine
- Set the mode
Your turn

Run the tree_spec chunk in your .qmd.
Edit this code to use a logistic regression model.
All available models are listed at https://www.tidymodels.org/find/parsnip/
Extension/Challenge: Edit this code to use a different model. For example, try using a conditional inference tree as implemented in the partykit package by changing the engine - or try an entirely different model type!
05:00
Models we’ll be using today
- Logistic regression
- Decision trees
Logistic regression
Logistic regression
Logistic regression
- Logit of outcome probability modeled as linear combination of predictors:
\(log(\frac{p}{1 - p}) = \beta_0 + \beta_1\cdot \text{A}\)
- Find a sigmoid line that separates the two classes
Decision trees
Decision trees
Series of splits or if/then statements based on predictors
First the tree grows until some condition is met (maximum depth, no more data)
Then the tree is pruned to reduce its complexity
Decision trees
All models are wrong, but some are useful!
Logistic regression
Decision trees
A model workflow
Workflows bind preprocessors and models
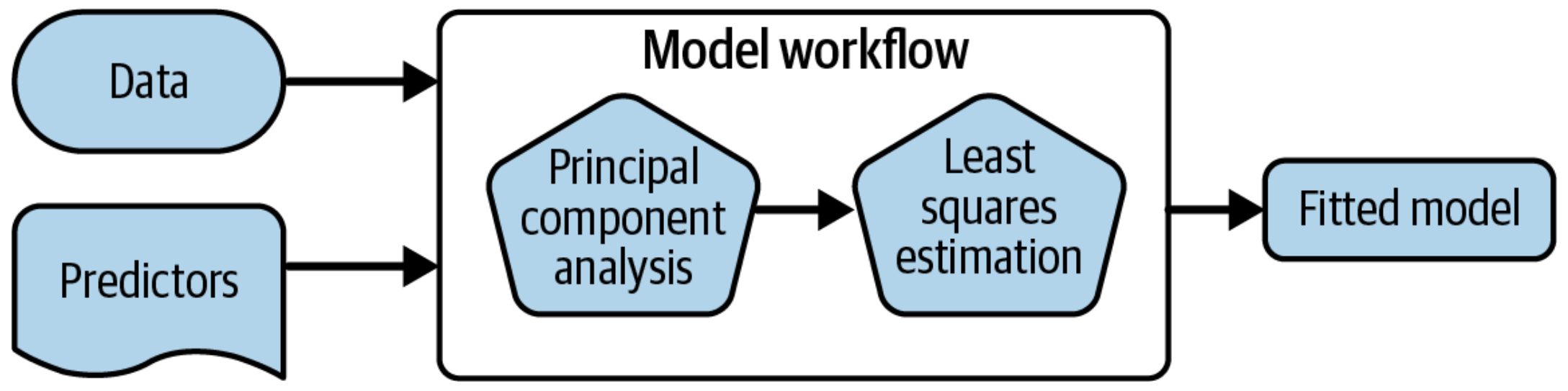
What is wrong with this?
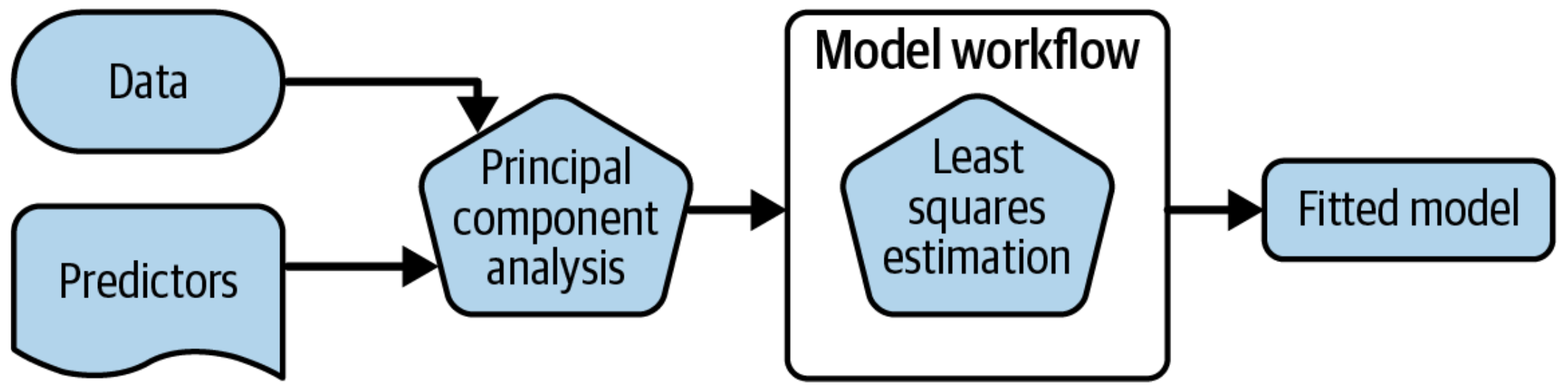
Why a workflow()? ![]()
- Workflows handle new data better than base R tools in terms of new factor levels
- You can use other preprocessors besides formulas (more on feature engineering in Advanced tidymodels!)
- They can help organize your work when working with multiple models
- Most importantly, a workflow captures the entire modeling process:
fit()andpredict()apply to the preprocessing steps in addition to the actual model fit
A model workflow ![]()
![]()
tree_spec <-
decision_tree() %>%
set_mode("classification")
tree_spec %>%
fit(forested ~ ., data = forested_train)
#> parsnip model object
#>
#> n= 5685
#>
#> node), split, n, loss, yval, (yprob)
#> * denotes terminal node
#>
#> 1) root 5685 2550 Yes (0.55145119 0.44854881)
#> 2) land_type=Tree 3064 300 Yes (0.90208877 0.09791123) *
#> 3) land_type=Barren,Non-tree vegetation 2621 371 No (0.14154903 0.85845097)
#> 6) temp_annual_max< 13.395 347 153 Yes (0.55907781 0.44092219)
#> 12) tree_no_tree=Tree 92 6 Yes (0.93478261 0.06521739) *
#> 13) tree_no_tree=No tree 255 108 No (0.42352941 0.57647059) *
#> 7) temp_annual_max>=13.395 2274 177 No (0.07783641 0.92216359) *A model workflow ![]()
![]()
tree_spec <-
decision_tree() %>%
set_mode("classification")
workflow() %>%
add_formula(forested ~ .) %>%
add_model(tree_spec) %>%
fit(data = forested_train)
#> ══ Workflow [trained] ════════════════════════════════════════════════
#> Preprocessor: Formula
#> Model: decision_tree()
#>
#> ── Preprocessor ──────────────────────────────────────────────────────
#> forested ~ .
#>
#> ── Model ─────────────────────────────────────────────────────────────
#> n= 5685
#>
#> node), split, n, loss, yval, (yprob)
#> * denotes terminal node
#>
#> 1) root 5685 2550 Yes (0.55145119 0.44854881)
#> 2) land_type=Tree 3064 300 Yes (0.90208877 0.09791123) *
#> 3) land_type=Barren,Non-tree vegetation 2621 371 No (0.14154903 0.85845097)
#> 6) temp_annual_max< 13.395 347 153 Yes (0.55907781 0.44092219)
#> 12) tree_no_tree=Tree 92 6 Yes (0.93478261 0.06521739) *
#> 13) tree_no_tree=No tree 255 108 No (0.42352941 0.57647059) *
#> 7) temp_annual_max>=13.395 2274 177 No (0.07783641 0.92216359) *A model workflow ![]()
![]()
tree_spec <-
decision_tree() %>%
set_mode("classification")
workflow(forested ~ ., tree_spec) %>%
fit(data = forested_train)
#> ══ Workflow [trained] ════════════════════════════════════════════════
#> Preprocessor: Formula
#> Model: decision_tree()
#>
#> ── Preprocessor ──────────────────────────────────────────────────────
#> forested ~ .
#>
#> ── Model ─────────────────────────────────────────────────────────────
#> n= 5685
#>
#> node), split, n, loss, yval, (yprob)
#> * denotes terminal node
#>
#> 1) root 5685 2550 Yes (0.55145119 0.44854881)
#> 2) land_type=Tree 3064 300 Yes (0.90208877 0.09791123) *
#> 3) land_type=Barren,Non-tree vegetation 2621 371 No (0.14154903 0.85845097)
#> 6) temp_annual_max< 13.395 347 153 Yes (0.55907781 0.44092219)
#> 12) tree_no_tree=Tree 92 6 Yes (0.93478261 0.06521739) *
#> 13) tree_no_tree=No tree 255 108 No (0.42352941 0.57647059) *
#> 7) temp_annual_max>=13.395 2274 177 No (0.07783641 0.92216359) *Your turn

Run the tree_wflow chunk in your .qmd.
Edit this code to make a workflow with your own model of choice.
Extension/Challenge: Other than formulas, what kinds of preprocessors are supported?
05:00
Predict with your model ![]()
![]()
How do you use your new tree_fit model?
Your turn

Run:
predict(tree_fit, new_data = forested_test)
What do you notice about the structure of the result?
03:00
Your turn

Run:
augment(tree_fit, new_data = forested_test)
How does the output compare to the output from predict()?
03:00
The tidymodels prediction guarantee!
- The predictions will always be inside a tibble
- The column names and types are unsurprising and predictable
- The number of rows in
new_dataand the output are the same
Understand your model ![]()
![]()
How do you understand your new tree_fit model?
Understand your model ![]()
![]()
How do you understand your new tree_fit model?
You can extract_*() several components of your fitted workflow.
⚠️ Never predict() with any extracted components!
Understand your model ![]()
![]()
How do you understand your new tree_fit model?
You can use your fitted workflow for model and/or prediction explanations:
- overall variable importance, such as with the vip package
- flexible model explainers, such as with the DALEXtra package
Learn more at https://www.tmwr.org/explain.html
Your turn

Extract the model engine object from your fitted workflow and check it out.
05:00
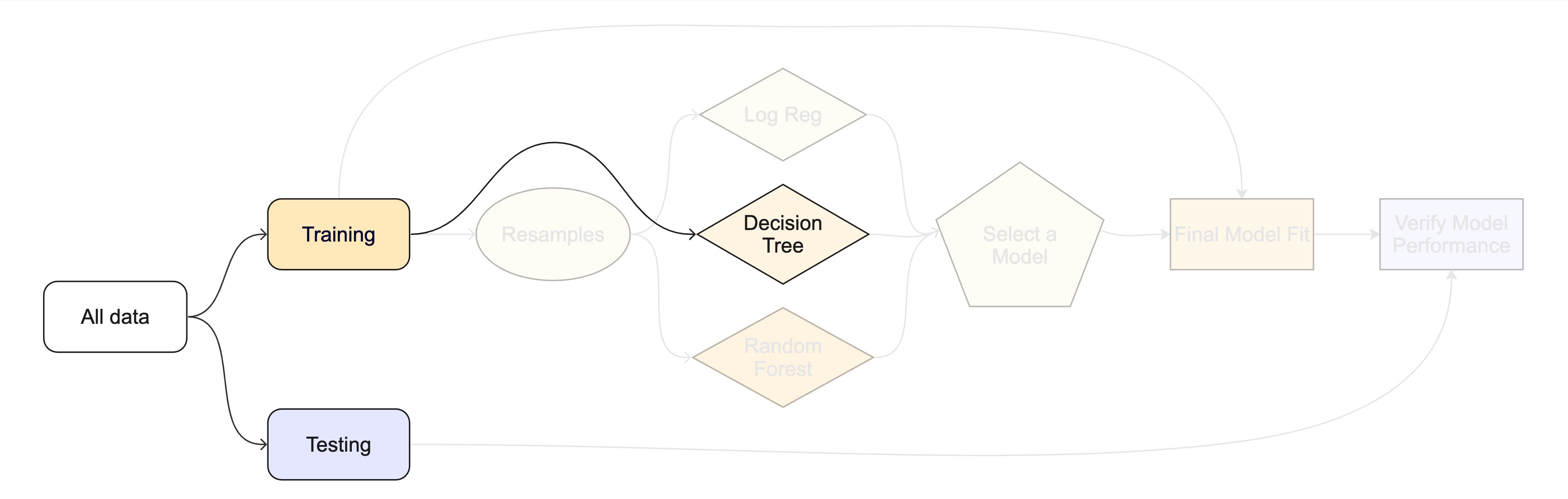
The whole game - status update